Exploratory Data Analysis#
# Basic System & Math Utilities
import os
import math
import glob
# Audio Processing Libraries
import wave
import librosa
import librosa.display
import soundfile
from scipy.io.wavfile import read
# Data Handling & Numerical Computing
import numpy as np
import pandas as pd
from scipy.stats import chi2_contingency
from scipy.stats import f_oneway
# Visualization Tools
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
# Machine Learning / Deep Learning
import tensorflow as tf
from tqdm.notebook import tqdm
# Jupyter Audio Playback
import IPython.display as ipd
from tqdm.notebook import tqdm
Explore Dataset Directory#
import kagglehub
vbookshelf_respiratory_sound_database_path = kagglehub.dataset_download('vbookshelf/respiratory-sound-database')
print('Data source import complete.')
Downloading from https://www.kaggle.com/api/v1/datasets/download/vbookshelf/respiratory-sound-database?dataset_version_number=2...
100%|██████████| 3.69G/3.69G [00:41<00:00, 95.2MB/s]
Extracting files...
Data source import complete.
The dataset used in this study is the Respiratory Sound Database, originally acquired from the Biomedical Health Informatics Challenge hosted by Aristotle University of Thessaloniki (Rocha et al., 2019). This database was curated by research teams from Portugal and Greece and is publicly accessible via Kaggle.
The dataset comprises 920 annotated audio recordings of respiratory sounds from 126 patients, with clips ranging from 10 to 90 seconds in length. Each recording includes corresponding annotation files that describe the presence of respiratory anomalies such as wheezes, crackles, or both, as well as demographic and diagnostic information for each patient. The dataset offers a comprehensive repository of real-world respiratory sound data, suitable for training and evaluating machine learning models targeting respiratory disease detection.
os.listdir(vbookshelf_respiratory_sound_database_path)
['respiratory_sound_database',
'Respiratory_Sound_Database',
'demographic_info.txt']
Load and Merge Demographics & Diagnosis CSVs#
df_no_diagnosis = pd.read_csv(
os.path.join(vbookshelf_respiratory_sound_database_path, 'demographic_info.txt'),
names=['Patient number', 'Age', 'Sex', 'Adult BMI (kg/m2)',
'Child Weight (kg)', 'Child Height (cm)'],
delimiter=' '
)
diagnosis = pd.read_csv(
os.path.join(vbookshelf_respiratory_sound_database_path, 'Respiratory_Sound_Database',
'Respiratory_Sound_Database', 'patient_diagnosis.csv'),
names=['Patient number', 'Diagnosis']
)
# Merge demographics + diagnosis
df = df_no_diagnosis.join(
diagnosis.set_index('Patient number'),
on='Patient number',
how='left'
)
df
| Patient number | Age | Sex | Adult BMI (kg/m2) | Child Weight (kg) | Child Height (cm) | Diagnosis | |
|---|---|---|---|---|---|---|---|
| 0 | 101 | 3.00 | F | NaN | 19.0 | 99.0 | URTI |
| 1 | 102 | 0.75 | F | NaN | 9.8 | 73.0 | Healthy |
| 2 | 103 | 70.00 | F | 33.00 | NaN | NaN | Asthma |
| 3 | 104 | 70.00 | F | 28.47 | NaN | NaN | COPD |
| 4 | 105 | 7.00 | F | NaN | 32.0 | 135.0 | URTI |
| ... | ... | ... | ... | ... | ... | ... | ... |
| 121 | 222 | 60.00 | M | NaN | NaN | NaN | COPD |
| 122 | 223 | NaN | NaN | NaN | NaN | NaN | COPD |
| 123 | 224 | 10.00 | F | NaN | 32.3 | 143.0 | Healthy |
| 124 | 225 | 0.83 | M | NaN | 7.8 | 74.0 | Healthy |
| 125 | 226 | 4.00 | M | NaN | 16.7 | 103.0 | Pneumonia |
126 rows × 7 columns
List All Filenames#
root = os.path.join(vbookshelf_respiratory_sound_database_path, 'Respiratory_Sound_Database', 'Respiratory_Sound_Database', 'audio_and_txt_files') + os.sep
filenames = [s.split('.')[0] for s in os.listdir(root) if s.endswith('.txt')]
Function to Extract Annotation Data#
def Extract_Annotation_Data(file_name, root):
tokens = file_name.split('_')
recording_info = pd.DataFrame(
data=[tokens],
columns=['Patient number', 'Recording index',
'Chest location', 'Acquisition mode', 'Recording equipment']
)
recording_annotations = pd.read_csv(
os.path.join(root, file_name + '.txt'),
names=['Start', 'End', 'Crackles', 'Wheezes'],
delimiter='\t'
)
return recording_info, recording_annotations
Collect Recording Info & Annotation Dictionaries#
i_list = []
rec_annotations_dict = {}
for s in filenames:
info, ann = Extract_Annotation_Data(s, root)
i_list.append(info)
rec_annotations_dict[s] = ann
recording_info = pd.concat(i_list, axis=0)
recording_info.tail()
| Patient number | Recording index | Chest location | Acquisition mode | Recording equipment | |
|---|---|---|---|---|---|
| 0 | 205 | 1b3 | Al | mc | AKGC417L |
| 0 | 166 | 1p1 | Pl | sc | Meditron |
| 0 | 109 | 1b1 | Al | sc | Litt3200 |
| 0 | 108 | 1b1 | Al | sc | Meditron |
| 0 | 159 | 1b1 | Al | sc | Meditron |
Count Crackles/Wheezes Per File#
filename_list = []
no_label_list = []
crack_list = []
wheeze_list = []
both_sym_list = []
for f in filenames:
d = rec_annotations_dict[f]
no_label = len(d[(d['Crackles'] == 0) & (d['Wheezes'] == 0)])
crack_only = len(d[(d['Crackles'] == 1) & (d['Wheezes'] == 0)])
wheeze_only = len(d[(d['Crackles'] == 0) & (d['Wheezes'] == 1)])
both = len(d[(d['Crackles'] == 1) & (d['Wheezes'] == 1)])
filename_list.append(f)
no_label_list.append(no_label)
crack_list.append(crack_only)
wheeze_list.append(wheeze_only)
both_sym_list.append(both)
file_label_df = pd.DataFrame({
'filename': filename_list,
'no label': no_label_list,
'crackles only': crack_list,
'wheezes only': wheeze_list,
'crackles and wheezees': both_sym_list
})
Aggregate & Assign Diagnosis Labels#
labels = file_label_df.sort_values(by='filename')
summary = labels.groupby('filename').sum()
conditions = [
(summary['crackles only'] == 0) &
(summary['wheezes only'] == 0) &
(summary['crackles and wheezees'] == 0),
(summary['crackles only'] == summary.max(axis=1)),
(summary['wheezes only'] == summary.max(axis=1)),
(summary['crackles and wheezees'] == summary.max(axis=1)),
(summary['no label'] == summary.max(axis=1)) &
(summary['crackles only'] > summary['wheezes only']) &
(summary['crackles only'] > summary['crackles and wheezees']),
(summary['no label'] == summary.max(axis=1)) &
(summary['wheezes only'] >= summary['crackles only']) &
(summary['wheezes only'] > summary['crackles and wheezees']),
(summary['no label'] == summary.max(axis=1)) &
(summary['crackles and wheezees'] >= summary['crackles only']) &
(summary['crackles and wheezees'] >= summary['wheezes only']),
]
values = [
'Healthy',
'Crackles',
'Wheezes',
'Wheezes & Crackles',
'Crackles',
'Wheezes',
'Wheezes & Crackles'
]
summary['diagnosis'] = np.select(conditions, values, default='Unknown')
summary
| no label | crackles only | wheezes only | crackles and wheezees | diagnosis | |
|---|---|---|---|---|---|
| filename | |||||
| 101_1b1_Al_sc_Meditron | 12 | 0 | 0 | 0 | Healthy |
| 101_1b1_Pr_sc_Meditron | 11 | 0 | 0 | 0 | Healthy |
| 102_1b1_Ar_sc_Meditron | 13 | 0 | 0 | 0 | Healthy |
| 103_2b2_Ar_mc_LittC2SE | 2 | 0 | 4 | 0 | Wheezes |
| 104_1b1_Al_sc_Litt3200 | 6 | 0 | 0 | 0 | Healthy |
| ... | ... | ... | ... | ... | ... |
| 224_1b2_Al_sc_Meditron | 7 | 0 | 0 | 0 | Healthy |
| 225_1b1_Pl_sc_Meditron | 14 | 0 | 0 | 0 | Healthy |
| 226_1b1_Al_sc_Meditron | 8 | 2 | 0 | 0 | Crackles |
| 226_1b1_Ll_sc_Meditron | 4 | 6 | 0 | 0 | Crackles |
| 226_1b1_Pl_sc_LittC2SE | 6 | 5 | 0 | 0 | Crackles |
920 rows × 5 columns
Exploratory Data Analysis#
Basic Overview of Demographic Dataset#
print("Shape of demographics dataframe:", df.shape)
df.head()
Shape of demographics dataframe: (126, 7)
| Patient number | Age | Sex | Adult BMI (kg/m2) | Child Weight (kg) | Child Height (cm) | Diagnosis | |
|---|---|---|---|---|---|---|---|
| 0 | 101 | 3.00 | F | NaN | 19.0 | 99.0 | URTI |
| 1 | 102 | 0.75 | F | NaN | 9.8 | 73.0 | Healthy |
| 2 | 103 | 70.00 | F | 33.00 | NaN | NaN | Asthma |
| 3 | 104 | 70.00 | F | 28.47 | NaN | NaN | COPD |
| 4 | 105 | 7.00 | F | NaN | 32.0 | 135.0 | URTI |
# Check missing values
df.isnull().sum()
| 0 | |
|---|---|
| Patient number | 0 |
| Age | 1 |
| Sex | 1 |
| Adult BMI (kg/m2) | 51 |
| Child Weight (kg) | 82 |
| Child Height (cm) | 84 |
| Diagnosis | 0 |
# Summary statistics
df.describe(include='all')
| Patient number | Age | Sex | Adult BMI (kg/m2) | Child Weight (kg) | Child Height (cm) | Diagnosis | |
|---|---|---|---|---|---|---|---|
| count | 126.000000 | 125.00000 | 125 | 75.000000 | 44.000000 | 42.000000 | 126 |
| unique | NaN | NaN | 2 | NaN | NaN | NaN | 8 |
| top | NaN | NaN | M | NaN | NaN | NaN | COPD |
| freq | NaN | NaN | 79 | NaN | NaN | NaN | 64 |
| mean | 163.500000 | 42.99264 | NaN | 27.190000 | 21.361136 | 104.652381 | NaN |
| std | 36.517119 | 32.20907 | NaN | 5.372519 | 17.150885 | 30.793128 | NaN |
| min | 101.000000 | 0.25000 | NaN | 16.500000 | 7.140000 | 64.000000 | NaN |
| 25% | 132.250000 | 4.00000 | NaN | 24.150000 | 11.755000 | 81.250000 | NaN |
| 50% | 163.500000 | 60.00000 | NaN | 27.400000 | 15.100000 | 99.500000 | NaN |
| 75% | 194.750000 | 71.00000 | NaN | 29.185000 | 24.325000 | 117.750000 | NaN |
| max | 226.000000 | 93.00000 | NaN | 53.500000 | 80.000000 | 183.000000 | NaN |
Diagnosis Distribution#
plt.figure(figsize=(8,5))
sns.countplot(data=df, x='Diagnosis')
plt.title("Distribution of Patient Diagnoses")
plt.xticks(rotation=45)
plt.show()
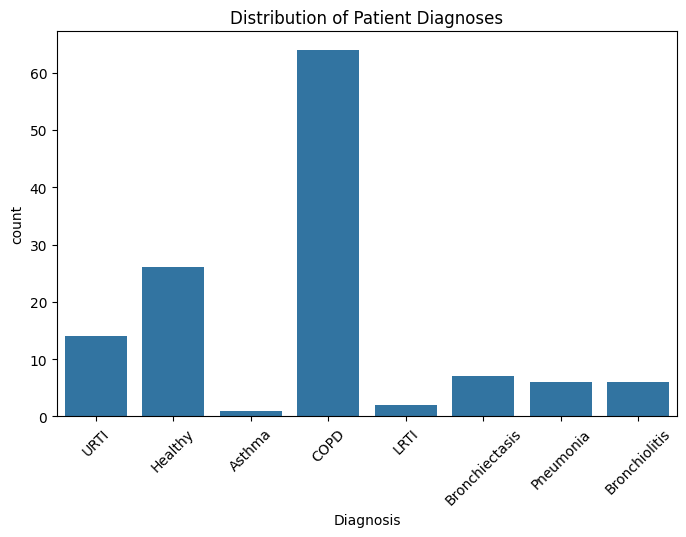
This bar chart shows the distribution of patient diagnoses in the dataset by count.
Dominant and Minority Classes#
COPD (Chronic Obstructive Pulmonary Disease) is the most represented diagnosis by far, with over 60 patients, making it the majority class in this sample.
Healthy individuals form the second largest group, with just over 25 cases, providing a sizeable control population for comparative analysis.
URTI (Upper Respiratory Tract Infection) is also notable, but substantially fewer than COPD and Healthy, with around 15 cases.
Diagnoses like Asthma, LRTI (Lower Respiratory Tract Infection), Bronchiectasis, Pneumonia, and Bronchiolitis are minority classes, each with less than 10 cases, indicating limited sampling or rarity in the dataset.
Analytical Implications#
The diagnosis distribution is highly imbalanced, especially with the dominance of COPD and the scarcity of less common conditions.
Models trained on this dataset may require strategies to handle class imbalance, such as oversampling minority classes, using class weights, or designing specific evaluation metrics to prevent bias toward COPD detection.
The strong representation of Healthy and COPD is beneficial for discriminative modeling but highlights the risk of reduced sensitivity and lower generalizability for rare diseases (e.g., Asthma, Bronchiolitis).
Epidemiological Considerations#
High COPD counts may reflect real-world prevalence patterns, especially in older, smoking, or at-risk populations, or could be due to targeted recruitment in the dataset.
The rarity of asthma and select infections could indicate these conditions are less prevalent in the sampled cohort or might result from inclusion criteria and annotation emphasis.
plt.figure(figsize=(7,5))
sns.countplot(data=df, x='Sex', hue='Diagnosis')
plt.title("Gender vs Diagnosis")
plt.show()
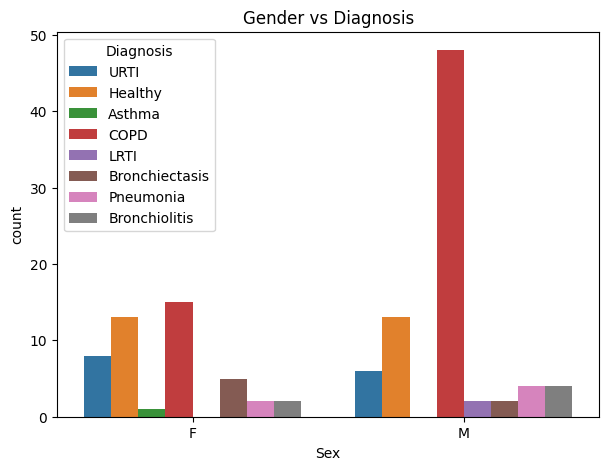
This grouped bar chart displays counts of recordings by diagnosis and sex (female “F” and male “M”) for several respiratory conditions.
Major Trends#
COPD (Chronic Obstructive Pulmonary Disease) shows a strong male predominance, with more than triple the number of male cases compared to females.
Healthy individuals are distributed equally between sexes, suggesting the reference population is balanced.
URTI (Upper Respiratory Tract Infection), Bronchiectasis, Pneumonia, and Bronchiolitis have lower and fairly balanced counts across both sexes, indicating no strong gender bias in sample collection for these conditions.
Observations#
Asthma is notably rare in this subset, with only 1 female case and none in males—highlighting potential sampling or annotation limitations for this diagnosis.
LRTI (Lower Respiratory Tract Infection) is more frequent in males but remains one of the less common classes overall.
Overview of Audio-Based Labels#
labels_df_recreated = file_label_df.sort_values(by='filename')
summary = labels_df_recreated.groupby('filename').sum()
# Re-apply the logic
conditions = [
(summary['crackles only'] == 0) &
(summary['wheezes only'] == 0) &
(summary['crackles and wheezees'] == 0),
(summary['crackles only'] == summary.max(axis=1)),
(summary['wheezes only'] == summary.max(axis=1)),
(summary['crackles and wheezees'] == summary.max(axis=1)),
(summary['no label'] == summary.max(axis=1)) &
(summary['crackles only'] > summary['wheezes only']) &
(summary['crackles only'] > summary['crackles and wheezees']),
(summary['no label'] == summary.max(axis=1)) &
(summary['wheezes only'] >= summary['crackles only']) &
(summary['wheezes only'] > summary['crackles and wheezees']),
(summary['no label'] == summary.max(axis=1)) &
(summary['crackles and wheezees'] >= summary['crackles only']) &
(summary['crackles and wheezees'] >= summary['wheezes only']),
]
values = [
'Healthy',
'Crackles',
'Wheezes',
'Wheezes & Crackles',
'Crackles',
'Wheezes',
'Wheezes & Crackles'
]
summary['diagnosis'] = np.select(conditions, values, default='Unknown')
# Reset the index to make 'filename' a regular column, which is often easier for merging.
# This ensures the actual filenames are in a column named 'filename'.
summary = summary.reset_index().rename(columns={'index': 'filename'})
# Ensure the 'filename' column is explicitly string type to use .str accessor reliably
summary['filename'] = summary['filename'].astype(str)
# Extract Patient number from the 'filename' column
# The filename format is 'PATIENT_NUMBER_RECORDING_INDEX_...'
summary['Patient number'] = summary['filename'].str.split('_').str[0].astype(int)
# Rename the existing audio-based diagnosis column before merging to avoid name collision
summary = summary.rename(columns={'diagnosis': 'audio_diagnosis'})
# Merge with the main demographic and diagnosis DataFrame (df)
# Select only the 'Patient number' and 'Diagnosis' columns from df
# Assuming 'df' is available from previous cells
df_patient_diagnosis = df[['Patient number', 'Diagnosis']].drop_duplicates(subset=['Patient number'])
# Perform the merge. The clinical diagnosis column will be named 'Diagnosis'.
summary = pd.merge(summary, df_patient_diagnosis, on='Patient number', how='left')
# Rename the merged clinical 'Diagnosis' column for clarity
summary = summary.rename(columns={'Diagnosis': 'clinical_diagnosis'})
summary.head()
| filename | no label | crackles only | wheezes only | crackles and wheezees | audio_diagnosis | Patient number | clinical_diagnosis | |
|---|---|---|---|---|---|---|---|---|
| 0 | 101_1b1_Al_sc_Meditron | 12 | 0 | 0 | 0 | Healthy | 101 | URTI |
| 1 | 101_1b1_Pr_sc_Meditron | 11 | 0 | 0 | 0 | Healthy | 101 | URTI |
| 2 | 102_1b1_Ar_sc_Meditron | 13 | 0 | 0 | 0 | Healthy | 102 | Healthy |
| 3 | 103_2b2_Ar_mc_LittC2SE | 2 | 0 | 4 | 0 | Wheezes | 103 | Asthma |
| 4 | 104_1b1_Al_sc_Litt3200 | 6 | 0 | 0 | 0 | Healthy | 104 | COPD |
Overview of Audio VS Demographic Datasets#
columns_to_check = ['Age', 'Sex', 'Diagnosis', 'audio_diagnosis']
# Extract 'Patient number' from the 'filename' column of the 'summary' DataFrame
summary_with_patient_id = summary.copy()
summary_with_patient_id['Patient number'] = summary_with_patient_id['filename'].str.split('_').str[0].astype(int)
# Merge df (demographic/clinical diagnosis) with summary (audio-based diagnosis)
# Only select the patient number and the audio-based diagnosis from summary_with_patient_id
merged_data = df.merge(summary_with_patient_id[['Patient number', 'audio_diagnosis']], on='Patient number', how='inner')
cleaned_merged_data = merged_data.dropna(subset=columns_to_check).copy()
print(f"Original merged_data shape: {merged_data.shape}")
print(f"Cleaned merged_data shape: {cleaned_merged_data.shape}")
print("First 5 rows of cleaned_merged_data:")
cleaned_merged_data.head()
Original merged_data shape: (920, 8)
Cleaned merged_data shape: (914, 8)
First 5 rows of cleaned_merged_data:
| Patient number | Age | Sex | Adult BMI (kg/m2) | Child Weight (kg) | Child Height (cm) | Diagnosis | audio_diagnosis | |
|---|---|---|---|---|---|---|---|---|
| 0 | 101 | 3.00 | F | NaN | 19.0 | 99.0 | URTI | Healthy |
| 1 | 101 | 3.00 | F | NaN | 19.0 | 99.0 | URTI | Healthy |
| 2 | 102 | 0.75 | F | NaN | 9.8 | 73.0 | Healthy | Healthy |
| 3 | 103 | 70.00 | F | 33.00 | NaN | NaN | Asthma | Wheezes |
| 4 | 104 | 70.00 | F | 28.47 | NaN | NaN | COPD | Healthy |
##Analyze Age vs. Respiratory Sound Patterns (ANOVA)
# Get unique values from the 'diagnosis' column
relevant_audio_diagnoses = cleaned_merged_data['audio_diagnosis'].unique()
# Create a list of age arrays for each audio-based diagnosis group
age_groups_audio = [cleaned_merged_data['Age'][cleaned_merged_data['audio_diagnosis'] == d].dropna() for d in relevant_audio_diagnoses]
# Perform one-way ANOVA test
f_statistic_audio, p_value_audio = f_oneway(*age_groups_audio)
# Calculate degrees of freedom
df_between_audio = len(relevant_audio_diagnoses) - 1
df_within_audio = sum(len(group) for group in age_groups_audio) - len(relevant_audio_diagnoses)
# Calculate eta-squared (η²) and eta (η)
eta_squared_audio = (f_statistic_audio * df_between_audio) / (f_statistic_audio * df_between_audio + df_within_audio)
eta_age_audio = np.sqrt(eta_squared_audio)
print(f"One-way ANOVA for Age vs. Respiratory Sound Patterns:")
print(f"F-statistic: {f_statistic_audio:.3f}")
print(f"P-value: {p_value_audio:.3f}")
print(f"Correlation Ratio (η) for Age vs. Respiratory Sound Patterns: {eta_age_audio:.3f}")
One-way ANOVA for Age vs. Respiratory Sound Patterns:
F-statistic: 17.229
P-value: 0.000
Correlation Ratio (η) for Age vs. Respiratory Sound Patterns: 0.232
One-way ANOVA shows a statistically significant difference in mean age across audio-based diagnosis groups (F(2, N−3) = 17.23, p < 0.001). However, the correlation ratio (η = 0.232; η² ≈ 0.054) indicates a small effect — only ~5.4% of age variance is explained by diagnosis group. Clinically, older participants appear slightly more likely to have crackles or mixed findings, but age alone is a weak predictor of respiratory sound class and should not be used as a surrogate for diagnostic category.
Analyze Gender vs. Respiratory Sound Patterns (Chi-squared)#
# 1. Create a contingency table
contingency_table_audio = pd.crosstab(cleaned_merged_data['Sex'], cleaned_merged_data['audio_diagnosis'])
# 3. Perform the Chi-squared test of independence
chi2_audio, p_audio, dof_audio, expected_audio = chi2_contingency(contingency_table_audio)
# 4. Print the chi-squared statistic and the p-value
print(f"Chi-squared statistic for Gender vs. Respiratory Sound Patterns: {chi2_audio:.3f}")
print(f"P-value for Gender vs. Respiratory Sound Patterns: {p_audio:.3f}")
Chi-squared statistic for Gender vs. Respiratory Sound Patterns: 25.522
P-value for Gender vs. Respiratory Sound Patterns: 0.000
def calculate_cramers_v(contingency_table, chi2):
n = contingency_table.sum().sum()
min_dim = min(contingency_table.shape)
v = np.sqrt(chi2 / (n * (min_dim - 1)))
return v
cramers_v_audio = calculate_cramers_v(contingency_table_audio, chi2_audio)
print(f"Cramers V for Gender vs. Respiratory Sound Patterns: {cramers_v_audio:.3f}")
Cramers V for Gender vs. Respiratory Sound Patterns: 0.167
The Chi-squared test showed a statistically significant association between gender and audio-based diagnosis (χ² = 25.52, p < 0.001). However, the effect size was small (Cramer’s V = 0.167), indicating that gender has only a weak relationship with the presence of crackles, wheezes, or combined sounds. Gender should therefore not be considered a strong predictor of respiratory sound category.
Analyze Age vs. Clinical Diagnosis (ANOVA)#
# 1. Get unique values from the 'Diagnosis' column
demographic_diagnoses = cleaned_merged_data['Diagnosis'].unique()
# 2. Create a list of age arrays for each demographic diagnosis group
age_groups_demographic = [cleaned_merged_data['Age'][cleaned_merged_data['Diagnosis'] == d].dropna() for d in demographic_diagnoses]
# 3. Perform one-way ANOVA test
f_statistic_demographic, p_value_demographic = f_oneway(*age_groups_demographic)
# Calculate degrees of freedom
df_between_demographic = len(demographic_diagnoses) - 1
df_within_demographic = sum(len(group) for group in age_groups_demographic) - len(demographic_diagnoses)
# Calculate eta-squared (η²) and eta (η)
eta_squared_demographic = (f_statistic_demographic * df_between_demographic) / (f_statistic_demographic * df_between_demographic + df_within_demographic)
eta_age_demographic = np.sqrt(eta_squared_demographic)
# 4. Print the F-statistic and p-value
print(f"One-way ANOVA for Age vs. Clinical Diagnoses:")
print(f"F-statistic: {f_statistic_demographic:.3f}")
print(f"P-value: {p_value_demographic:.3f}")
print(f"Correlation Ratio (η) for Age vs. Clinical Diagnoses: {eta_age_demographic:.3f}")
One-way ANOVA for Age vs. Clinical Diagnoses:
F-statistic: 418.525
P-value: 0.000
Correlation Ratio (η) for Age vs. Clinical Diagnoses: 0.874
One-way ANOVA revealed a highly significant difference in age across demographic diagnosis groups (F = 418.53, p < 0.001). The correlation ratio was very large(η = 0.874, η² ≈ 0.763), indicating that diagnosis category strongly depends on age. Approximately 76% of the variance in age is explained by diagnostic group assignment, reflecting the well-known age-related distribution of diseases such as COPD (older adults) versus asthma (younger individuals).
Analyze Gender vs. Clinical Diagnosis (Chi-squared)#
# 1. Create a contingency table
contingency_table_demographic = pd.crosstab(cleaned_merged_data['Sex'], cleaned_merged_data['Diagnosis'])
# 2. Perform the Chi-squared test of independence
chi2_demographic, p_demographic, dof_demographic, expected_demographic = chi2_contingency(contingency_table_demographic)
# 3. Print the chi-squared statistic and the p-value
print(f"Chi-squared statistic for Gender vs. Clinical Diagnoses: {chi2_demographic:.3f}")
print(f"P-value for Gender vs. Clinical Diagnoses: {p_demographic:.3f}")
Chi-squared statistic for Gender vs. Clinical Diagnoses: 31.036
P-value for Gender vs. Clinical Diagnoses: 0.000
cramers_v_demographic = calculate_cramers_v(contingency_table_demographic, chi2_demographic)
print(f"Cramers V for Gender vs. Clinical Diagnoses: {cramers_v_demographic:.3f}")
Cramers V for Gender vs. Clinical Diagnoses: 0.184
This shows a statistically significant association between gender and respiratory diagnoses (p = 0.000), indicating that certain conditions are more common in one gender than the other. However, the strength of this association is weak to moderate (Cramér’s V = 0.184), meaning gender alone is not a strong predictor of diagnosis. Clinically, while slight gender trends may exist, gender should be considered alongside other features for accurate assessment or modeling.
Example Waveform and Spectrogram Visual#
i = 100
sound_filename = root + file_label_df['filename'][i] + '.wav'
ipd.Audio(sound_filename, rate=16000)
target_diagnoses = ['COPD', 'Healthy', 'URTI', 'Bronchiectasis', 'Pneumonia', 'Bronchiolitis']
# Define hop_length explicitly so both RMS and Time calculation use the same value
hop_length = 512
for diagnosis_type in target_diagnoses:
# Find the first filename for the current diagnosis type
# Note: Ensure 'summary' dataframe and 'root' path are defined in your environment
# The 'filename' column must be present in the 'summary' DataFrame after jo2rAMCoOT09 is run correctly.
example_filename = summary[summary['clinical_diagnosis'] == diagnosis_type]['filename'].iloc[0]
# Construct the full path to the WAV file
wav_path = os.path.join(root, example_filename + '.wav')
# Load the audio file
x, sr = librosa.load(wav_path, sr=16000)
# Calculate RMS energy
# Explicitly passing hop_length ensures consistency
rms = librosa.feature.rms(y=x, hop_length=hop_length)
# We must tell times_like what the SR and hop_length are
times = librosa.times_like(rms, sr=sr, hop_length=hop_length)
# Create a figure and axes for plotting
plt.figure(figsize=(12, 4))
ax = plt.subplot(1, 1, 1)
# Plot the waveform
librosa.display.waveshow(y=x, sr=sr, ax=ax, alpha=0.6)
# Overlay the RMS envelope
ax.plot(times, rms[0], color='r', linestyle='-', label='RMS Energy')
ax.legend()
# Set plot title
ax.set(title=f'Sample Waveform for {diagnosis_type}')
plt.tight_layout()
plt.show()
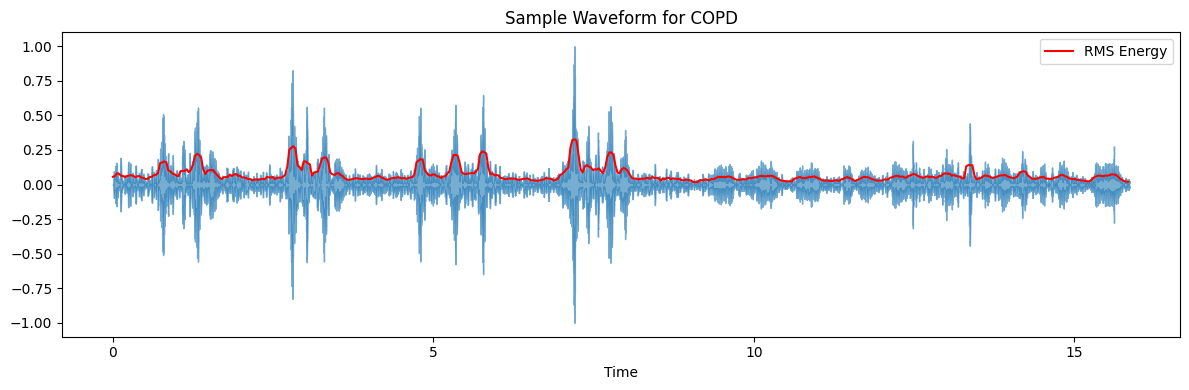
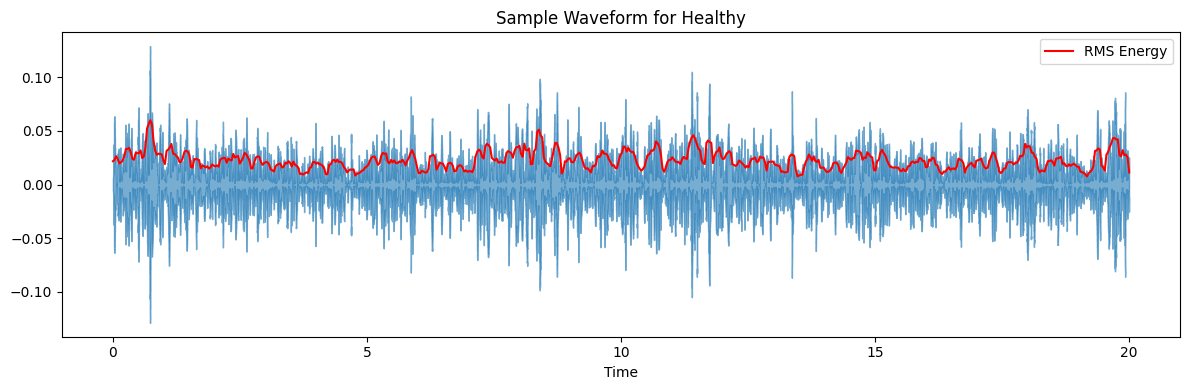
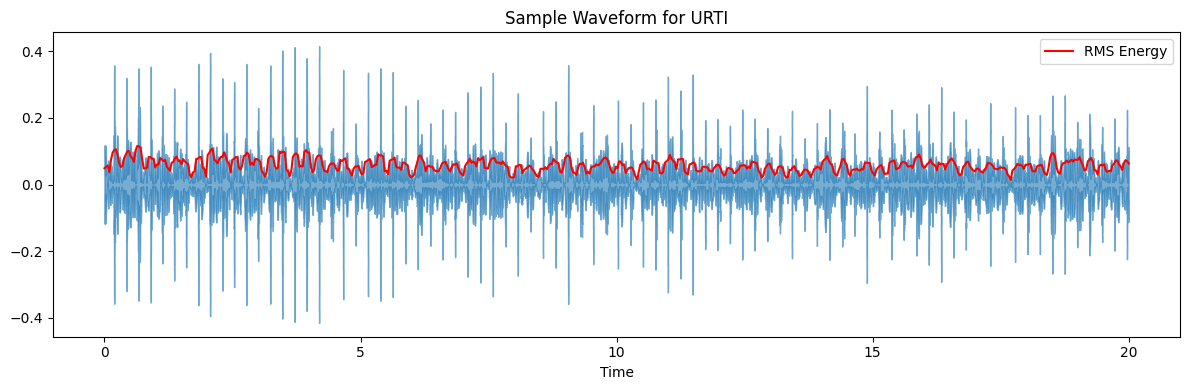
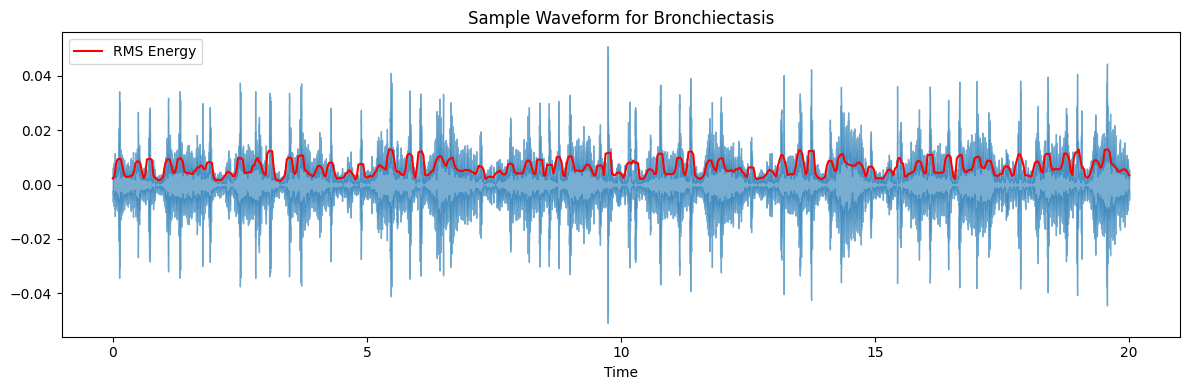
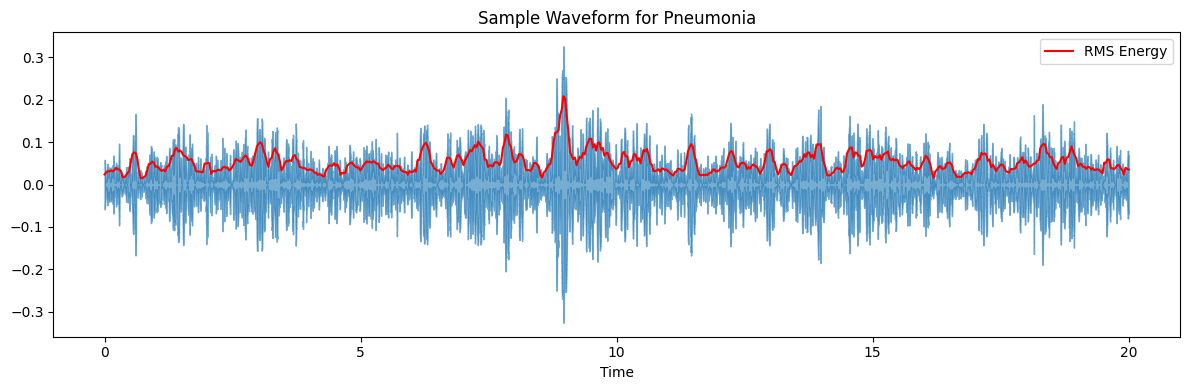
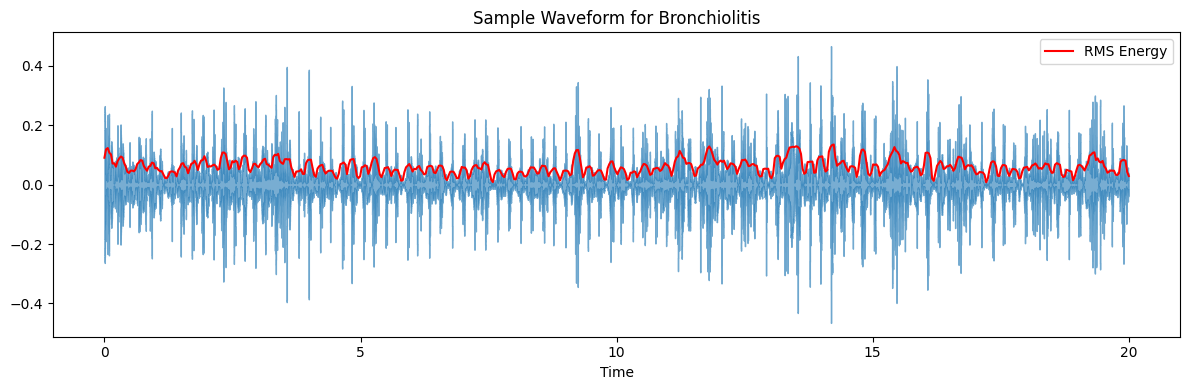
target_plot_categories = ['COPD', 'Healthy', 'Pneumonia', 'Others']
hop_length = 512
for category_type in target_plot_categories:
if category_type == 'Others':
# Select one diagnosis from the 'Others' group to represent it
# 'URTI' is chosen as a representative example from the 'Others' group
example_diagnosis_for_others = 'URTI'
example_filename = summary[summary['clinical_diagnosis'] == example_diagnosis_for_others]['filename'].iloc[0]
plot_title_diagnosis = f'Sample Waveform for {category_type} (e.g., {example_diagnosis_for_others})'
else:
example_filename = summary[summary['clinical_diagnosis'] == category_type]['filename'].iloc[0]
plot_title_diagnosis = f'Sample Waveform for {category_type}'
# Construct the full path to the WAV file
wav_path = os.path.join(root, example_filename + '.wav')
# Load the audio file
x, sr = librosa.load(wav_path, sr=16000)
# Calculate RMS energy
rms = librosa.feature.rms(y=x, hop_length=hop_length)
# Define times for RMS envelope
times = librosa.times_like(rms, sr=sr, hop_length=hop_length)
# Create a figure and axes for plotting
plt.figure(figsize=(12, 4))
ax = plt.subplot(1, 1, 1)
# Plot the waveform
librosa.display.waveshow(y=x, sr=sr, ax=ax, alpha=0.6)
# Overlay the RMS envelope
ax.plot(times, rms[0], color='r', linestyle='-', label='RMS Energy')
ax.legend()
# Set plot title
ax.set(title=plot_title_diagnosis)
plt.tight_layout()
plt.show()



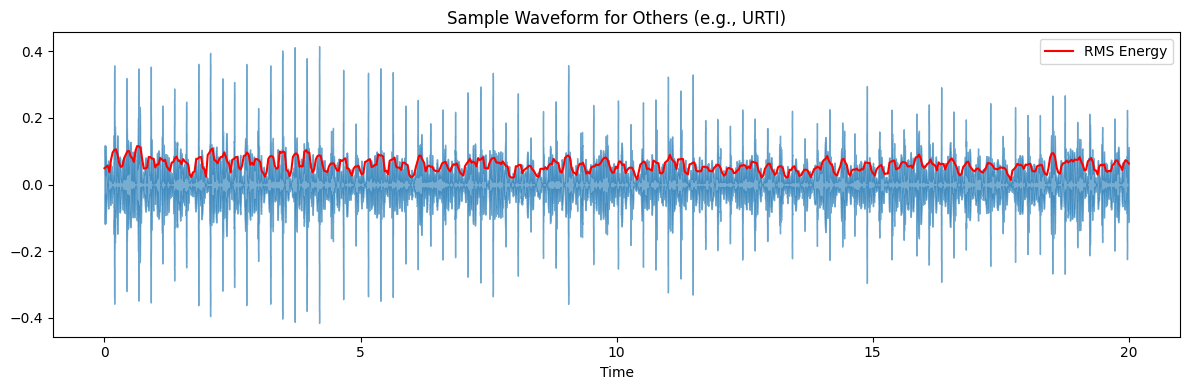
Extract Audio Features#
# 1. Re-create summary DataFrame with clinical diagnosis to ensure data consistency
labels_df_recreated = file_label_df.sort_values(by='filename')
summary_temp = labels_df_recreated.groupby('filename').sum()
conditions = [
(summary_temp['crackles only'] == 0) &
(summary_temp['wheezes only'] == 0) &
(summary_temp['crackles and wheezees'] == 0),
(summary_temp['crackles only'] == summary_temp.max(axis=1)),
(summary_temp['wheezes only'] == summary_temp.max(axis=1)),
(summary_temp['crackles and wheezees'] == summary_temp.max(axis=1)),
(summary_temp['no label'] == summary_temp.max(axis=1)) &
(summary_temp['crackles only'] > summary_temp['wheezes only']) &
(summary_temp['crackles only'] > summary_temp['crackles and wheezees']),
(summary_temp['no label'] == summary_temp.max(axis=1)) &
(summary_temp['wheezes only'] >= summary_temp['crackles only']) &
(summary_temp['wheezes only'] > summary_temp['crackles and wheezees']),
(summary_temp['no label'] == summary_temp.max(axis=1)) &
(summary_temp['crackles and wheezees'] >= summary_temp['crackles only']) &
(summary_temp['crackles and wheezees'] >= summary_temp['wheezes only']),
]
values = [
'Healthy',
'Crackles',
'Wheezes',
'Wheezes & Crackles',
'Crackles',
'Wheezes',
'Wheezes & Crackles'
]
summary_temp['diagnosis'] = np.select(conditions, values, default='Unknown')
# Reset the index to make 'filename' a regular column
summary_merged = summary_temp.reset_index().rename(columns={'index': 'filename'})
# Ensure the 'filename' column is explicitly string type
summary_merged['filename'] = summary_merged['filename'].astype(str)
# Extract Patient number from the 'filename' column
summary_merged['Patient number'] = summary_merged['filename'].str.split('_').str[0].astype(int)
# Rename the existing audio-based diagnosis column
summary_merged = summary_merged.rename(columns={'diagnosis': 'audio_diagnosis'})
# Merge with the main demographic and diagnosis DataFrame (df)
df_patient_diagnosis = df[['Patient number', 'Diagnosis']].drop_duplicates(subset=['Patient number'])
summary_merged = pd.merge(summary_merged, df_patient_diagnosis, on='Patient number', how='left')
# Rename the merged clinical 'Diagnosis' column for clarity
summary_merged = summary_merged.rename(columns={'Diagnosis': 'clinical_diagnosis'})
print("Recreated summary_merged DataFrame with clinical diagnosis.")
# 2. & 3. Extract Audio Features and create features_df
all_audio_features = []
sr_new = 16000 # Standardize sample rate to 16kHz for feature extraction
for index, row in tqdm(summary_merged.iterrows(), total=summary_merged.shape[0], desc="Extracting Audio Features"):
filename = row['filename']
clinical_diagnosis = row['clinical_diagnosis']
audio_diagnosis = row['audio_diagnosis']
wav_path = os.path.join(root, filename + '.wav')
try:
y, sr_original = librosa.load(wav_path, sr=None) # Load with original sample rate
# Resample if original sample rate is different from target
if sr_original != sr_new:
y_resampled = librosa.resample(y=y, orig_sr=sr_original, target_sr=sr_new)
else:
y_resampled = y
# Calculate duration
duration = len(y_resampled) / sr_new # Use resampled length and new SR
# Calculate Spectral Centroid
spectral_centroid = np.mean(librosa.feature.spectral_centroid(y=y_resampled, sr=sr_new))
# Calculate Spectral Bandwidth
spectral_bandwidth = np.mean(librosa.feature.spectral_bandwidth(y=y_resampled, sr=sr_new))
# Calculate Spectral Roll-off
spectral_rolloff = np.mean(librosa.feature.spectral_rolloff(y=y_resampled, sr=sr_new))
# Calculate Spectral Flux using onset_strength as a proxy
onset_env = librosa.onset.onset_strength(y=y_resampled, sr=sr_new)
spectral_flux = np.mean(onset_env)
all_audio_features.append({
'filename': filename,
'clinical_diagnosis': clinical_diagnosis,
'audio_diagnosis': audio_diagnosis,
'original_sample_rate': sr_original,
'duration_sec': duration,
'spectral_centroid': spectral_centroid,
'spectral_bandwidth': spectral_bandwidth,
'spectral_rolloff': spectral_rolloff,
'spectral_flux': spectral_flux
})
except Exception as e:
print(f"Error processing {filename}: {e}")
all_audio_features.append({
'filename': filename,
'clinical_diagnosis': clinical_diagnosis,
'audio_diagnosis': audio_diagnosis,
'original_sample_rate': None,
'duration_sec': None,
'spectral_centroid': None,
'spectral_bandwidth': None,
'spectral_rolloff': None,
'spectral_flux': None
})
features_df = pd.DataFrame(all_audio_features)
print("Audio feature extraction complete.")
# 4. Group by clinical_diagnosis and aggregate, creating a new 'grouped_clinical_diagnosis'
def group_diagnoses_for_mean_features(diagnosis):
if diagnosis in ['Healthy', 'COPD', 'Pneumonia']:
return diagnosis
else:
return 'Others'
features_df['grouped_clinical_diagnosis'] = features_df['clinical_diagnosis'].apply(group_diagnoses_for_mean_features)
mean_features_by_grouped_clinical_diagnosis = features_df.groupby('grouped_clinical_diagnosis').agg(
**{
'Mean Sample Rate (Hz)': ('original_sample_rate', 'mean'),
'Mean Duration (s)': ('duration_sec', 'mean'),
'Min Duration (s)': ('duration_sec', 'min'),
'Max Duration (s)': ('duration_sec', 'max'),
'Mean Spectral Centroid (Hz)': ('spectral_centroid', 'mean'),
'Mean Spectral Bandwidth (Hz)': ('spectral_bandwidth', 'mean'),
'Mean Spectral Rolloff (Hz)': ('spectral_rolloff', 'mean'),
'Mean Spectral Flux (Hz)': ('spectral_flux', 'mean')
}
)
# 5. Filter for requested diagnoses
target_display_diagnoses = ['Healthy', 'COPD', 'Pneumonia', 'Others']
# Handle cases where a diagnosis might not be present in the grouped results
filtered_mean_features = mean_features_by_grouped_clinical_diagnosis.reindex(target_display_diagnoses).copy()
Recreated summary_merged DataFrame with clinical diagnosis.
Audio feature extraction complete.
display(filtered_mean_features)
| Mean Sample Rate (Hz) | Mean Duration (s) | Min Duration (s) | Max Duration (s) | Mean Spectral Centroid (Hz) | Mean Spectral Bandwidth (Hz) | Mean Spectral Rolloff (Hz) | Mean Spectral Flux (Hz) | |
|---|---|---|---|---|---|---|---|---|
| grouped_clinical_diagnosis | ||||||||
| Healthy | 44100.000000 | 19.986857 | 19.800 | 20.000000 | 209.567730 | 719.838010 | 219.702667 | 0.777009 |
| COPD | 39290.920555 | 21.732459 | 7.856 | 86.200000 | 353.635311 | 884.412032 | 519.429352 | 0.834581 |
| Pneumonia | 44100.000000 | 20.000000 | 20.000 | 20.000000 | 138.338795 | 565.168151 | 131.423843 | 0.540922 |
| Others | 44100.000000 | 19.993655 | 19.820 | 20.000062 | 229.009911 | 655.560376 | 282.665291 | 0.821843 |
features_to_plot = [
'spectral_centroid',
'spectral_bandwidth',
'spectral_rolloff',
'spectral_flux'
]
# For display purposes in the title, we can map the internal column names to more readable names
feature_display_names = {
'spectral_centroid': 'Spectral Centroid (Hz)',
'spectral_bandwidth': 'Spectral Bandwidth (Hz)',
'spectral_rolloff': 'Spectral Rolloff (Hz)',
'spectral_flux': 'Spectral Flux (Hz)'
}
# Create a new 'grouped_diagnosis' column
def group_diagnoses(diagnosis):
if diagnosis in ['Healthy', 'COPD', 'Pneumonia']:
return diagnosis
else:
return 'Others'
features_df_grouped = features_df.copy()
features_df_grouped['grouped_diagnosis'] = features_df_grouped['clinical_diagnosis'].apply(group_diagnoses)
plt.figure(figsize=(20, 16))
for i, feature_col_name in enumerate(features_to_plot):
plt.subplot(2, 2, i + 1) # 2 rows, 2 columns
sns.boxplot(
data=features_df_grouped,
x='grouped_diagnosis', # Use the new grouped diagnosis column
y=feature_col_name, # Use the actual column name
palette='viridis'
)
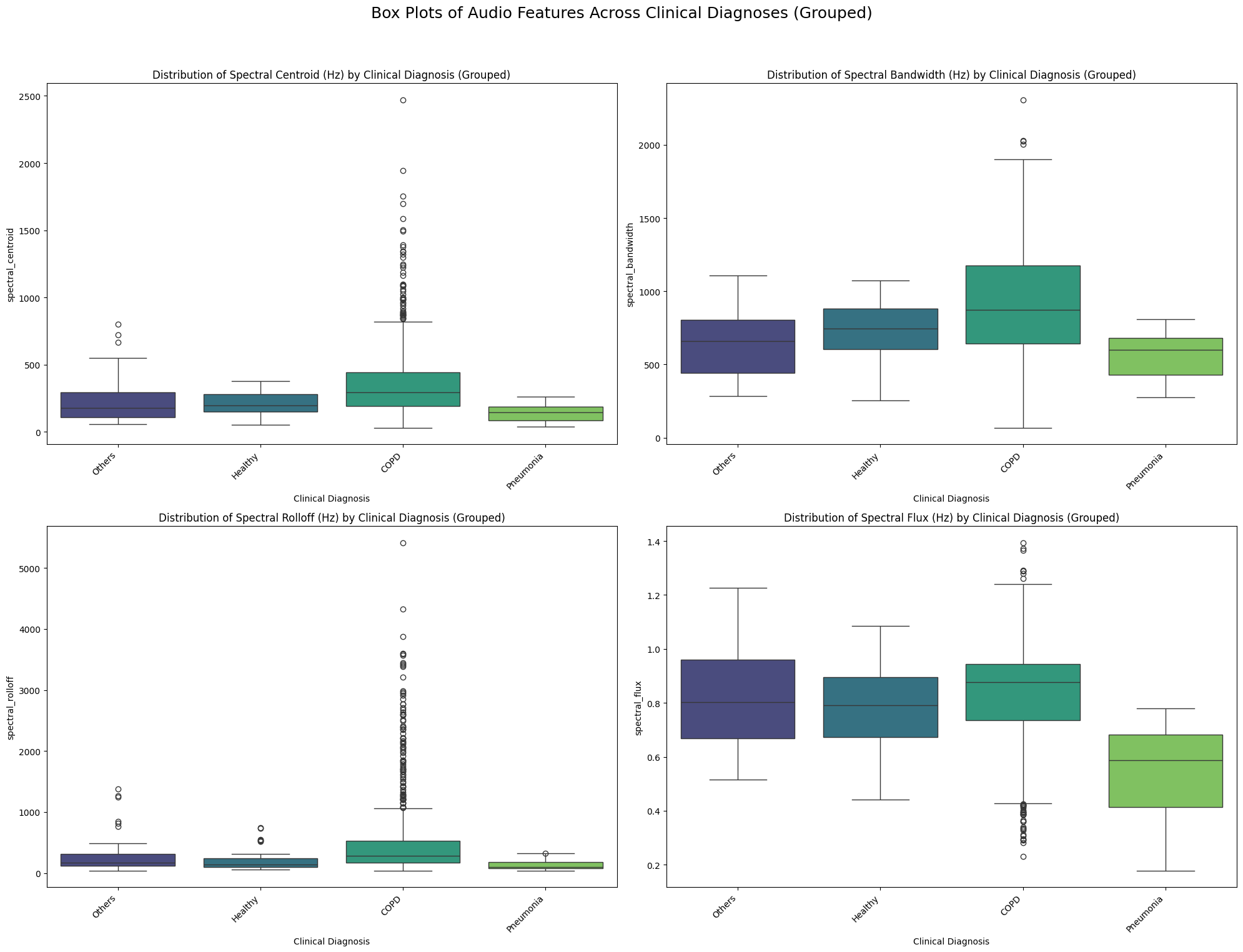
plt.title(f'Distribution of {feature_display_names[feature_col_name]} by Clinical Diagnosis (Grouped)') # Use display name in title
plt.xlabel('Clinical Diagnosis') # Add a clear label for the x-axis
plt.xticks(rotation=45, ha='right')
plt.suptitle('Box Plots of Audio Features Across Clinical Diagnoses (Grouped)', fontsize=18)
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
plt.show()
/tmp/ipython-input-1969003633.py:30: FutureWarning:
Passing `palette` without assigning `hue` is deprecated and will be removed in v0.14.0. Assign the `x` variable to `hue` and set `legend=False` for the same effect.
sns.boxplot(
/tmp/ipython-input-1969003633.py:30: FutureWarning:
Passing `palette` without assigning `hue` is deprecated and will be removed in v0.14.0. Assign the `x` variable to `hue` and set `legend=False` for the same effect.
sns.boxplot(
/tmp/ipython-input-1969003633.py:30: FutureWarning:
Passing `palette` without assigning `hue` is deprecated and will be removed in v0.14.0. Assign the `x` variable to `hue` and set `legend=False` for the same effect.
sns.boxplot(
/tmp/ipython-input-1969003633.py:30: FutureWarning:
Passing `palette` without assigning `hue` is deprecated and will be removed in v0.14.0. Assign the `x` variable to `hue` and set `legend=False` for the same effect.
sns.boxplot(